Honey is a well-known natural food source, made by bees using nectar from flowers. Typically honey is classified by color and flavor depending on the type of flower from which the nectar was harvested from.
Pure honey is a complex supersaturated solution containing 80% sugars and unique combination of chemical compounds such as phenolic, flavonoids, and volatiles suspended in water. Every honey has antimicrobial properties due to hydrogen peroxide, which is a naturally present mild antiseptic. Hydrogen peroxide is known to be unstable and can be quickly destroyed in the presence of heat, light, and body enzymes which, reduces its medicinal properties.
Manuka honey is obtained from the nectar of the flower of the manuka bush (Leptospermum scoparium), which is native to New Zealand and some parts of Australia. It is widely recognised for its non-peroxide antimicrobial properties and specific taste. The chemical compounds that make Manuka honey so special are believed to be Methylglyoxal (MGO), Leptosperin and Dihydroxyacetone (DHA). Methylglyoxal is a naturally occurring dicarbonyl compound produced through the conversion of DHA during maturation and aging of honey and is believed to be the major contributor to manuka honey’s non-peroxide antibacterial activity. It undergoes a spontaneous reaction with water, and it has been estimated that in aqueous solution less than 1% of the MGO remains unreacted. It predominantly forms the two compounds methylglyoxal monohydrate and methylglyoxal dihydrate as shown in Fig. 1. Due to its reactivity, low molecular mass and lack of UV chromophore, the direct detection of MGO in manuka honey is problematic using conventional methods [1].
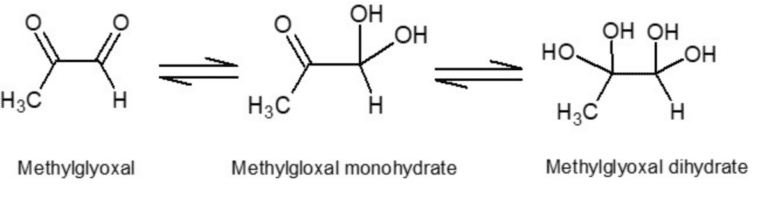
Figure 1: The structures of the MGO molecule and the two hydrates.
The quality of Manuka honey is specified by the UMF (Unique Manuka Factor) grading system, which represents the unique signature compounds characteristic of this honey, including Leptosperin, DHA and MGO. Higher UMF numbers stipulate a higher health benefit. For example, UMF-16 honey contains roughly 570 mg MGO per kg of dry honey [2].
In this work, we identified methylglyoxal hydrates and quantified them using an internal standard (maleic acid) with the help of benchtop NMR Spectroscopy. To perform this measurement, two sets of samples were prepared:
1. Methylglyoxal dissolved in a 100 mM Maleic Acid in D2O solution.
2. 100 mg manuka honey (UMF-16) dissolved in 1 mL of D2O .
0.5 mL of each sample was taken into three different standard 5 mm NMR tubes. The Proton spectra were collected on a Spinsolve 60 MHz benchtop NMR system using 256 scans, 15 s repetition time, with a total acquisition time of 16 minutes per spectrum. The Proton spectrum of sample 2 (UMF-16 manuka honey) is shown in Fig. 2. We can immediately identify the sugar peaks, but also smaller peaks at 7.4, 2.3 and 1.4 ppm that can be assigned to the unique manuka honey ingredients leptosperin, MGO mono- and dihydrate [3].
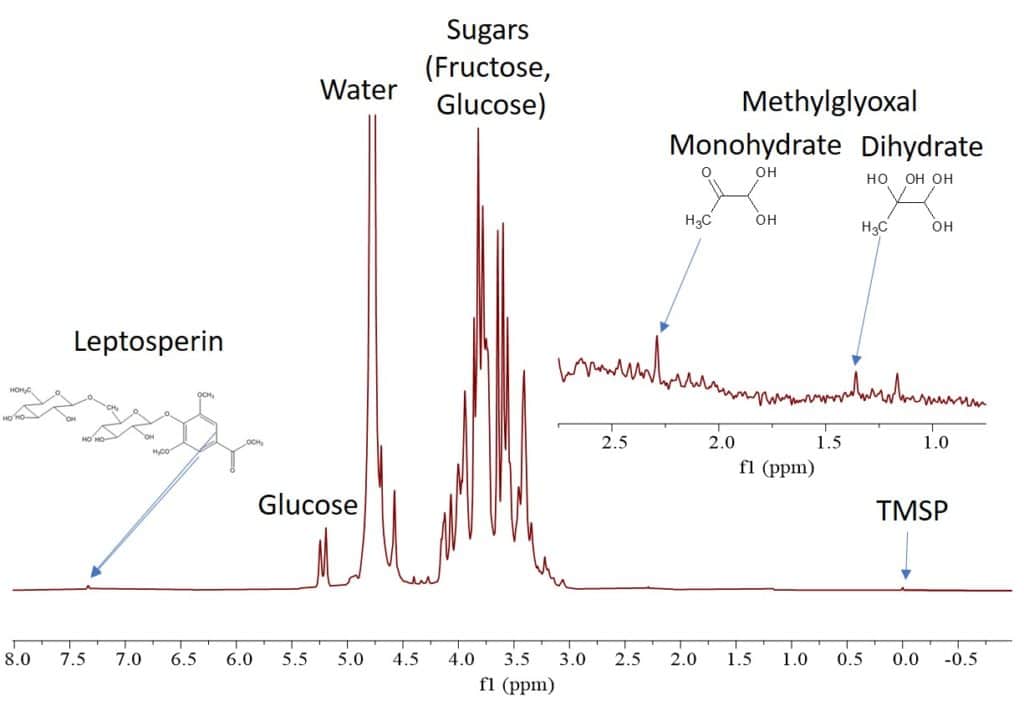
Figure 2: Proton NMR spectrum of UMF-16 manuka honey dissolved in D2O.
By using the qNMR method [4], we can quantify the MGO content in the UMF-16 manuka honey sample. For this we prepared sample 1 as external standard. Figure 3 shows the spectra of the two samples in the range of the MGO peaks.
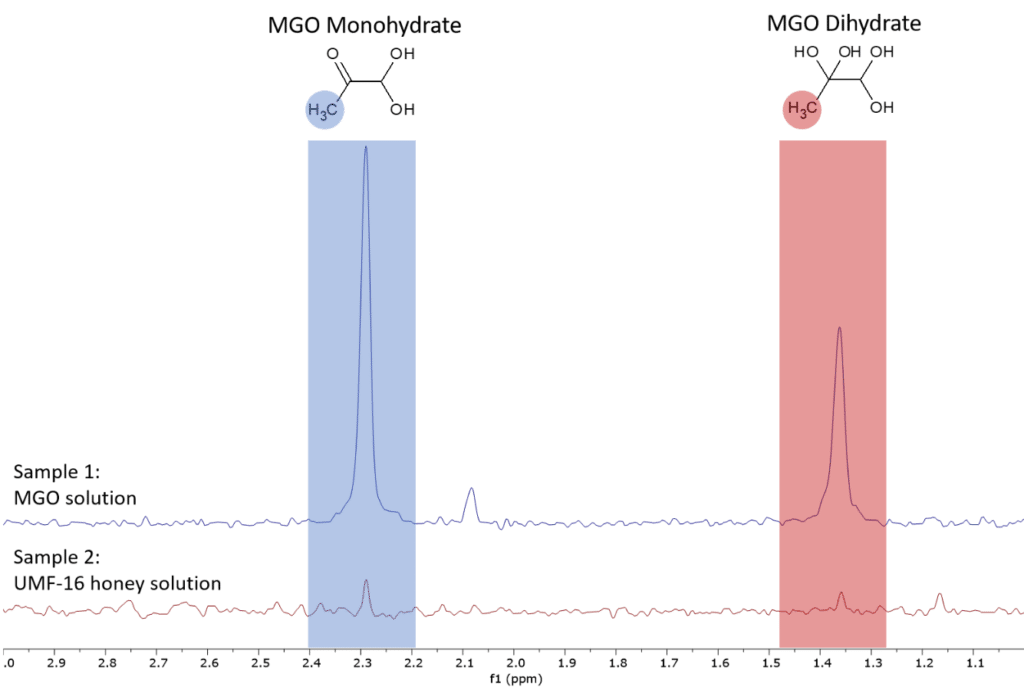
Figure 3: Proton NMR spectra of the two samples zoomed into range of the MGO peaks.
Sample 1 contained a precisely prepared amount of maleic acid as internal standard as specified in Table 1. By integrating the MGO peaks, we can calculate the MGO concentration of each sample. The total MGO concentration is listed in the last row of Table 1.
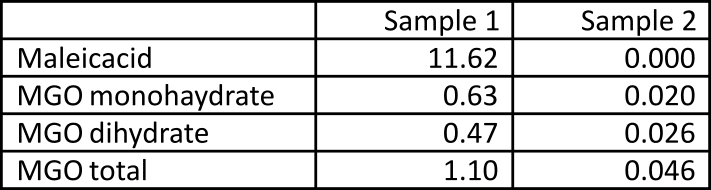
Table 1: Mass concentrations in mg/mL as obtained by qNMR of MGO in the two samples.
Sample 2 contains 100 mg of untreated manuka honey with a total of 46 ug MGO per mL. Considering that honey typically contains 20% of moisture, we have 80 mg of dried honey in the sample. Therefore the MGO concentration as measured by qNMR is 0.046/80*1000000=575 mg/kg. This measured value is in excellent agreement with the expected MGO concentration for UMF-16 manuka honey [2].
Please access the pdf version of the application note here.
References
[1] J. A. Donarski, D. P. T. Roberts, A. J. Charlton, Anal. Methods, 2010, 2, 1479-1483
[2] https://newzealandhoneyco.com/pages/mgo-vs-umf-calculator-manuka-honey
[3] M. Spiteri, K. M. Rogers, E. Jamin, F. Thomas, S. Guyader, M. Lees, D. N. Rutledge, Food Chem., 2017, 217, 766-772
[4] https://magritek.com/2016/09/12/quantitative-benchtop-nmr/